van start tot finish
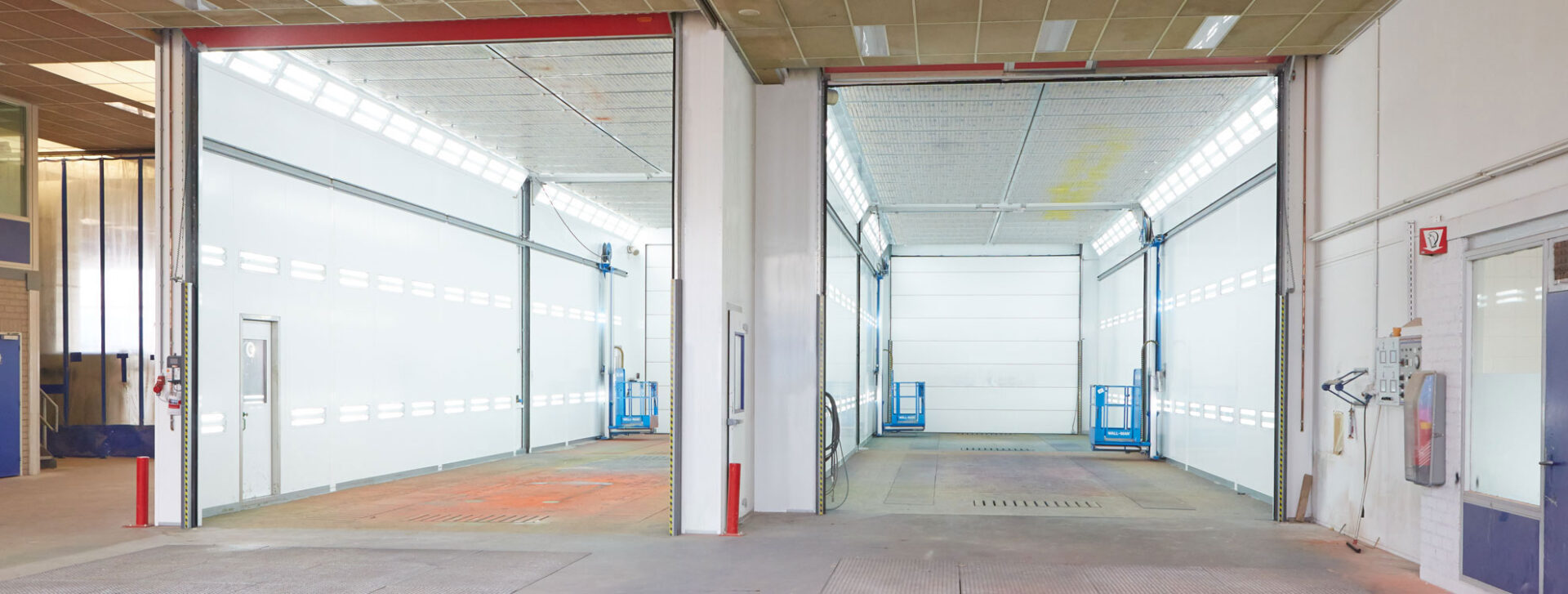
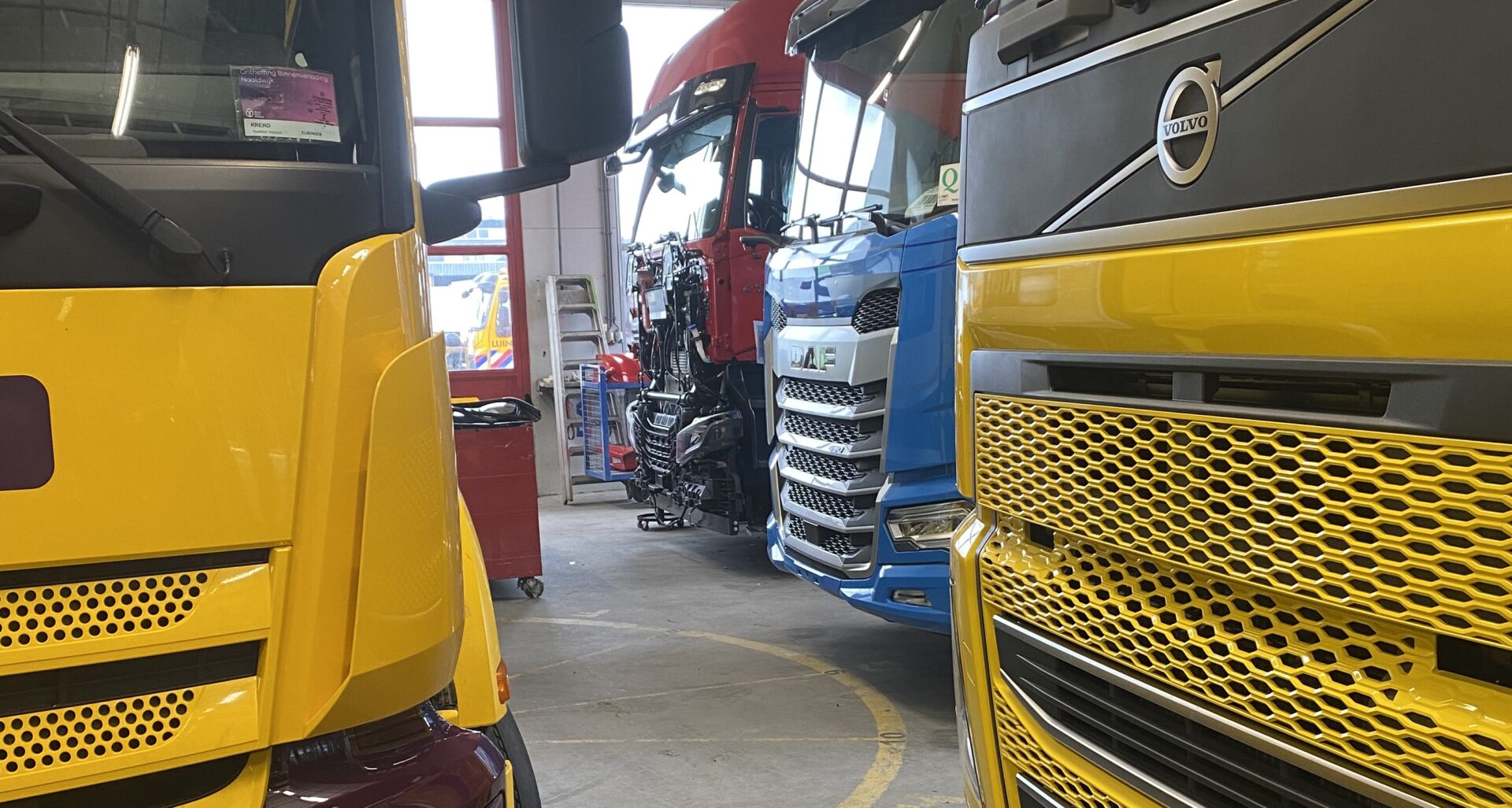
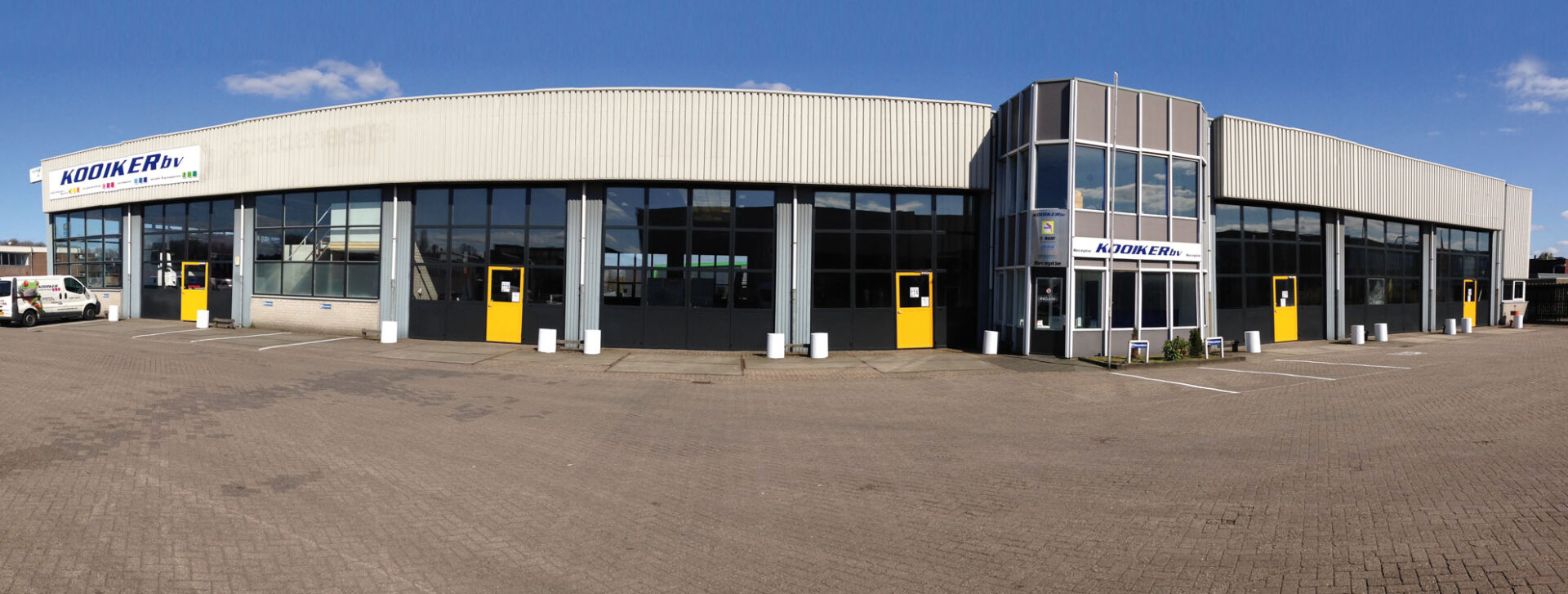
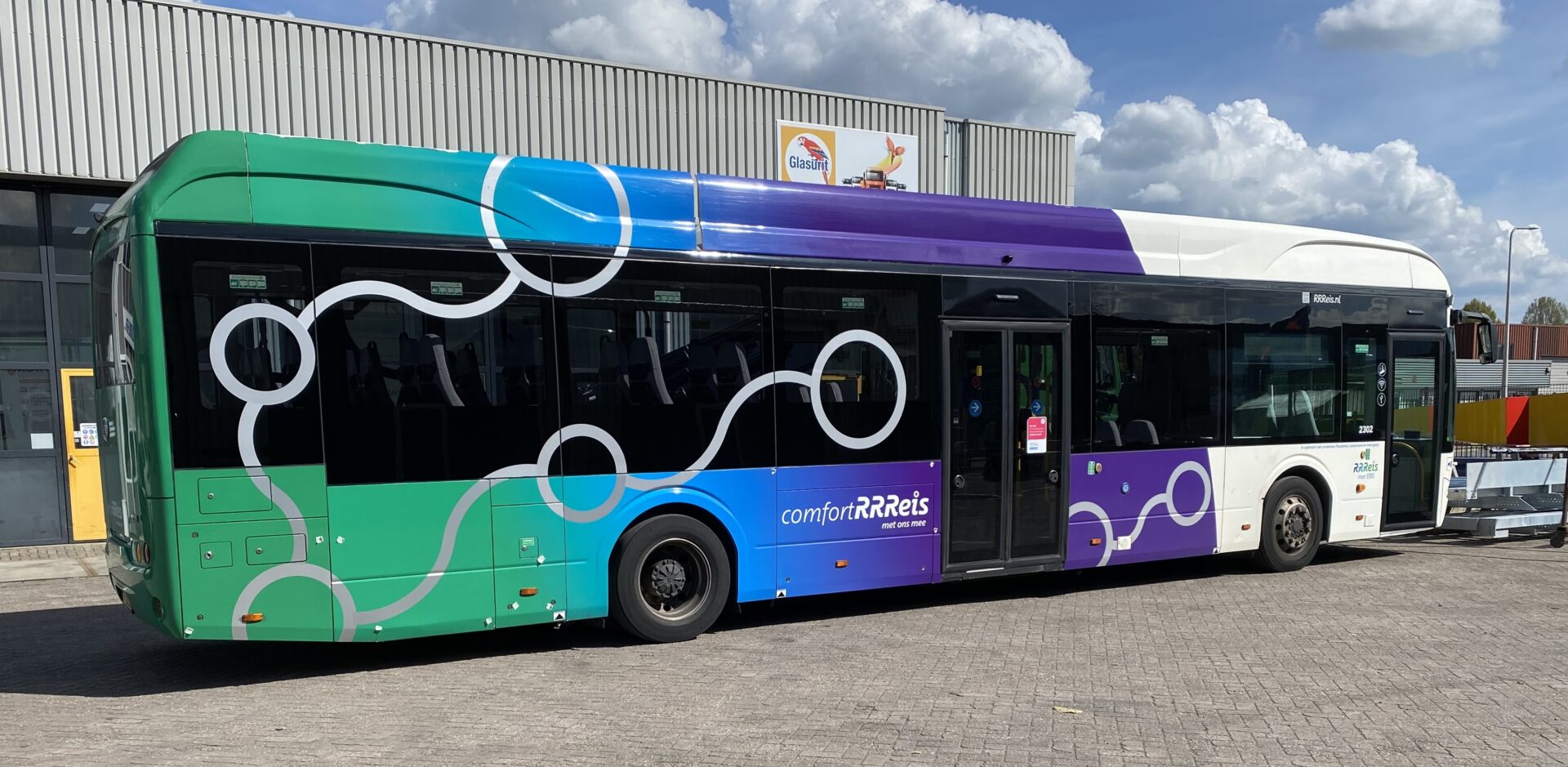
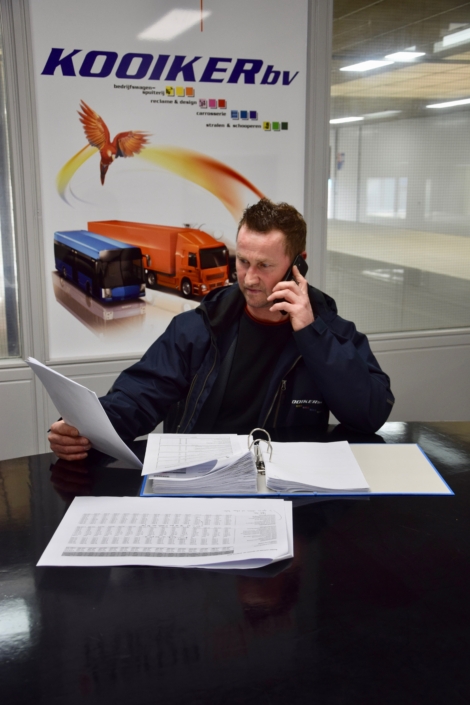
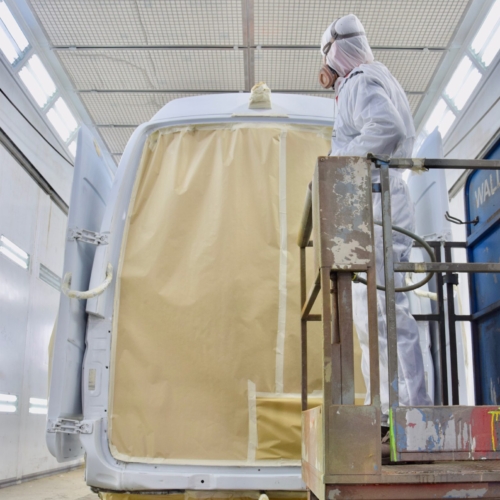
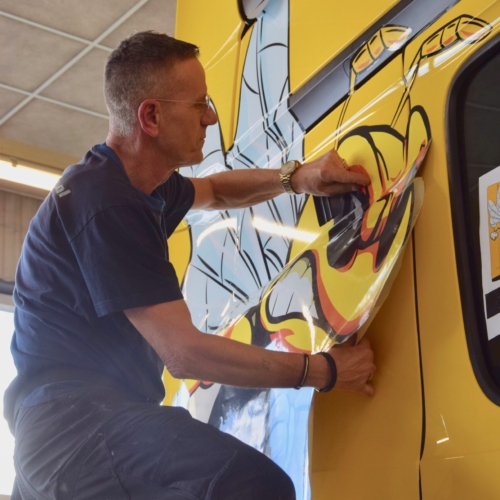
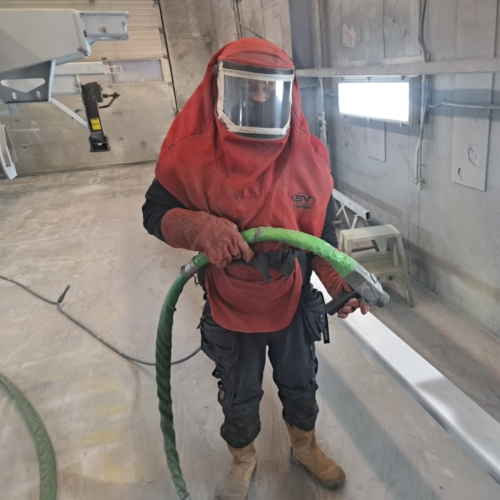
Uw bedrijfswagens zijn een visitekaartje van uw bedrijf. Dagelijks worden deze wagens door heel veel mensen op de meest uiteenlopende plaatsen gezien. Op de (snel)weg, in de dorpskern, bij het laden en lossen of op de parkeerplaats bij het benzinestation. Waarbij een herkenbaar logo of opvallende uitstraling ervoor zorgt dat klanten u herkennen en dat potentiële klanten u herinneren. Wij spuiten nieuwe of gebruikte bedrijfswagens in de door u gewenste huisstijl of voorzien die van reclame-uitingen. Alles op maat en alles onder één dak. Dus niet eerst naar andere locaties voor schadeherstel of laadklep. Eén locatie, één aanspreekpunt, één factuur. Dat spaart u tijd en dus kosten. En natuurlijk RAI CarrosserieNL gegarandeerde kwaliteit. Dus of u nu één wagen op de weg heeft of meer dan honderd, Kooiker biedt u een compleet dienstenpakket en is graag ook uw totaalpartner. Van start tot finish.